Filling in the Bird Tree of Life
[Posted by Chuck Almdale]
Note to readers: Over the past few months I have seen the diagram/painting below appear in many places. This circular “tree of life”-style diagram looks interesting and claims to include 10,135 species of birds, but the detail is far too small to read or be of use. I have searched unsuccessfully for an interactive version. If you find such a version, send me a link and I’ll post it. Lacking that, I include below links and snapshots of the OneZoom Trees of Life, which I blogged about in December 2015.
The tremendous changes in our understanding of phylogenetic relationships continue to mount. Since I began birding in the late 1970’s, almost 2,000 species have been added, some from field work but many from genetic analysis in the laboratory. The number of avian families has varied from 171 (Wetmore 1960) to 143 using a DNA-DNA hybridization technique (Sibley & Monroe, 1990 & 1993), to the current 249 based on the whole genome analysis (Cornell Lab Birds of the World, Apr 2021).
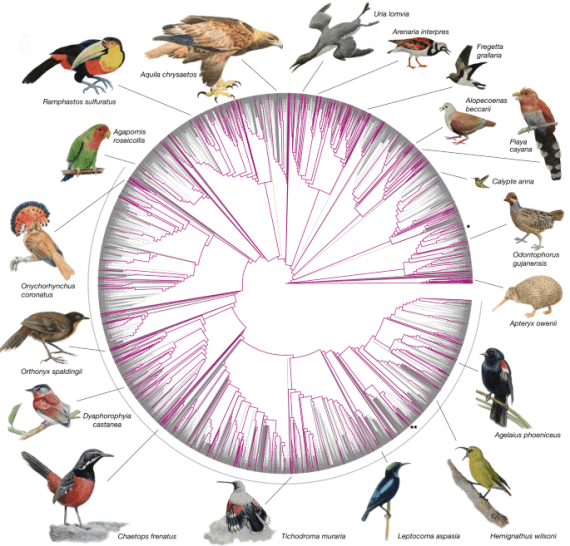
Fig. 1. 10,135 bird species are shown on a draft phylogeny that synthesizes taxonomic and phylogenetic information. In total, 363 species, covering 92.4% of all families, now have at least 1 genome assembly per sequenced family (purple branches). The grey arc (lower half) marks the diverse Passeriformes radiation, with 6,063 species, of which 173 species have genome assemblies now. Chicken (*) and zebra finch (**) are marked for orientation. Center point indicates the last common ancestor of all birds around 150 Ma (million years ago). Paintings illustrate examples of sequenced species. Image from Nature 11-Nov-2020, Dense sampling of bird diversity increases power of comparative genomics.
The chart above came from the following article.
Dense sampling of bird diversity increases power of comparative genomics.
Nature.com | Shaohong Feng, Josefin Stiller,…Guojie Zhang | 11 Nov 2020
The following is their abstract. The entire article is free to read on the link.
Abstract
Whole-genome sequencing projects are increasingly populating the tree of life and characterizing biodiversity. Sparse taxon sampling has previously been proposed to confound phylogenetic inference, and captures only a fraction of the genomic diversity. Here we report a substantial step towards the dense representation of avian phylogenetic and molecular diversity, by analysing 363 genomes from 92.4% of bird families—including 267 newly sequenced genomes produced for phase II of the Bird 10,000 Genomes (B10K) Project. We use this comparative genome dataset in combination with a pipeline that leverages a reference-free whole-genome alignment to identify orthologous regions in greater numbers than has previously been possible and to recognize genomic novelties in particular bird lineages. The densely sampled alignment provides a single-base-pair map of selection, has more than doubled the fraction of bases that are confidently predicted to be under conservation and reveals extensive patterns of weak selection in predominantly non-coding DNA. Our results demonstrate that increasing the diversity of genomes used in comparative studies can reveal more shared and lineage-specific variation, and improve the investigation of genomic characteristics. We anticipate that this genomic resource will offer new perspectives on evolutionary processes in cross-species comparative analyses and assist in efforts to conserve species.
The following chart is from an earlier study of 48 species representing the 40 orders of Neoaves from Struthioniformes (Ostrich) to three major divisions of Passeriformes (Perching or Song birds). This came from the following article.
Whole-genome analyses resolve early branches in the tree of life of modern birds.
ScienceMag.org | Jarvis, Mirarab, Aberer, Li, Houde, Ho, et.al. | 12 Dec 2014
There is a slightly interactive version of this chart here (it doubles in size).
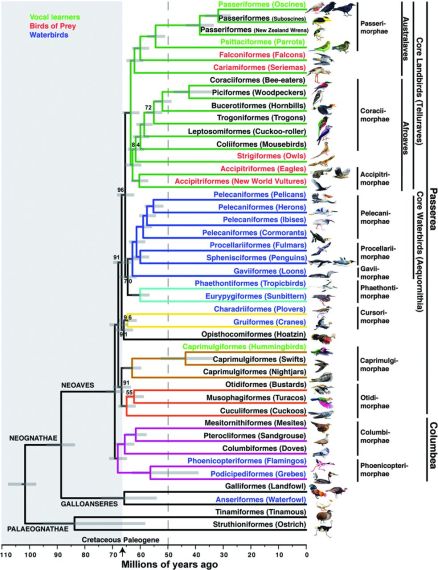
Fig. 1. Genome-scale phylogeny of birds.
Branch colors denote well-supported clades in this and other analyses. Names on branches denote orders (-iformes) and English group terms (in parentheses); drawings are of the specific species sequenced. To the right are superorder (-imorphae) and higher unranked names. In some groups, more than one species was sequenced, and these branches have been collapsed. Text color denotes groups of species with broadly shared traits, whether by homology or convergence. The arrow indicates the K-Pg boundary at 66 Ma, with the Cretaceous period shaded at left. The gray dashed line represents the approximate end time (50 Ma) by which nearly all neoavian orders diverged. Horizontal gray bars on each node indicate the 95% credible interval of divergence time in millions of years.
Finally, here’s a few screensnips from the OneZoom Trees of Life. These are completely interactive on their website, and you could spend the rest of your life looking up the relationships of everything of interest to you. Watch out!
Here’s their basic tree, covering earthly life from bacteria to eukaryotes. In case you’re wondering, you are an eukaryote, one of many.

A bit closer in, here’s the turtle-crocodile-bird branch.

If you want to look at just birds, this has 9,993 of them, no turtles need apply.

Let’s zoom in to our Northern Mockingbird. First we have to find the Passerines. As with using the alphabetically organized dictionary, it helps to have at least a vague idea where it might be located. Passerines (perching birds or song birds) are the most recently evolved avian order, so they’ll be at the farthest end of the curling branches, at the tip of that long one on the right.

Now that we found the Passerines, let’s zoom in some more and find Mockingbirds. There they are, buried among the Starlings. Didn’t know there were so many Starling species, did you? Looking at this, you can assume that Thrashers & Mockingbirds are — evolutionarily speaking — actually a divergent group of Starlings. How did this come about?
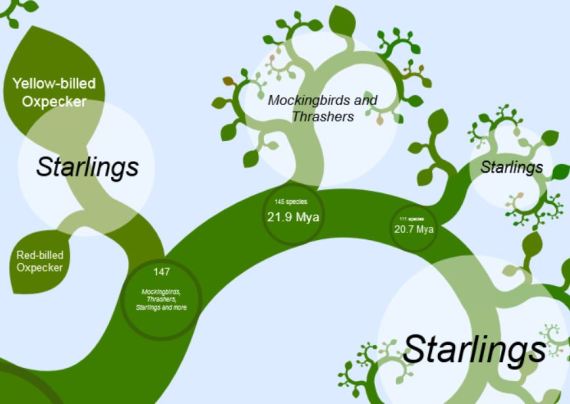
As the 34 species of Mockingbirds & Thrashers are all New World birds, and the remaining 113 species of Starlings and allies are all Old World Birds (never mind those European Starlings squeaking away outside your window, they’re human imports), it’s (relatively) safe to assume that somewhere around 23-28 million years ago (according to the little divergence dates in the Tree of Life) a few ancestral starlings flew over from Asia or Europe, settled in the New World, and speciated. That, in a nutshell, is one of the main features of evolution by means of natural selection.
And we finally get down to our Northern Mockingbird, whose closest relatives are the Socorro (splitting off 2.33 million years ago) and Tropical Mockingbird (split off 1.61 Mya).
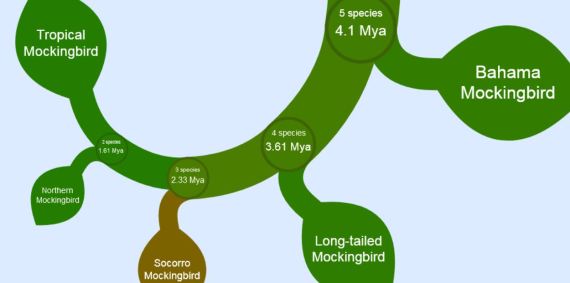
If you know exactly what you’re looking for, you can open the search bar, type in “Northern Mockingbird” click <Next Hit> and you’re there, without all the zooming in which might give your right index finger a repetitive stress injury. Then you can zoom back out.
I think this is a absolutely terrific program. There have been permanent links to it on our blogsite’s right-side sidebar (in “Other Blogs” near the bottom) for about five years.
Discover more from SANTA MONICA BAY AUDUBON SOCIETY BLOG
Subscribe to get the latest posts sent to your email.


